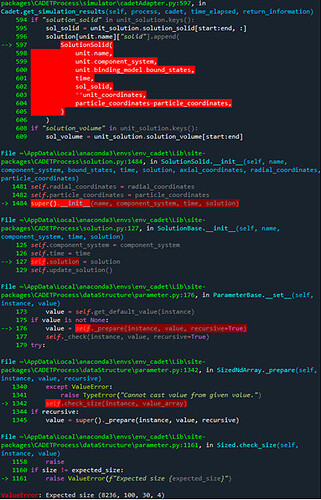
Here it is! Slightly delayed, but the problem still persists 
import numpy as np
import sys, glob
import pandas as pd
import matplotlib.pyplot as plt
from itertools import cycle
import time
from CADETProcess.processModel import ComponentSystem
from CADETProcess.processModel import MultiComponentColloidal
from CADETProcess.processModel import Inlet, GeneralRateModel, Outlet, GRMDiscretizationFV
from CADETProcess.processModel import FlowSheet
from CADETProcess.processModel import Process, solutionRecorder
from CADETProcess.processModel import LumpedRateModelWithoutPores
""" EXPERIMENTAL INPUTS _________________________________________________ """
MODEL_CADET = True
# Load description
VOL_LOAD = 12.5 #mL
C_LOAD = 12. # Load total concentration in g/L
HMW1_LOAD_PERCENT = 2.4 #2.4 # Small HMW (oligo)
HMW2_LOAD_PERCENT = 2.8 #3.5 # Large HMW
C_SALT = 100+17 # mM
# System description
COL_VOL = 0.7 # Column volume (mL)
DEAD_VOL = 0.25 #mL
# Process description
FLOWRATE = 0.18 # mL/min
COLVOL_WASH = 10
COLVOL_GRAD = 10
# UV-300 to concentration calibration curve: abs = a*conc
calibration_const_a = 40 # mAu / (mg/mL)
#Other
HMW_SCALE_FACTOR = 20
""" EXPERIMENTAL SECTION _________________________________________________ """
# Select chromatogram(s) to analyze by index
keep_idx = [0]
# Color cycle (matplotlib default)
color_list = plt.rcParams['axes.prop_cycle'].by_key()['color']
color_cycle = cycle(color_list)
# Create figure with twin axes
fig, ax1 = plt.subplots(figsize=(10,6))
ax2 = ax1.twinx()
# Gather CSV files and pick selected ones
files = sorted(glob.glob("*.csv"))
selected_files = [files[i] for i in keep_idx if 0 <= i < len(files)]
missing = [i for i in keep_idx if not (0 <= i < len(files))]
if missing:
print(f"Skipping out-of-range indices: {missing}")
""" *** CADET AND CHROMATOGRAPHY MODELING SECTION ************************* """
""" USER INPUTS ___________________________________________________________ """
mab_molecular_weight = 150 #kDa
load_conc = C_LOAD # g/L
column_length = 0.1 # m
column_dia = 0.003 # m
# Resin
column_porosity = 0.4
particle_porosity = 0.65
particle_diameter = 75E-6 # m
c_mab = 0.01*C_LOAD*(100-HMW1_LOAD_PERCENT-HMW2_LOAD_PERCENT)/mab_molecular_weight
c_hmw1 = 0.01*C_LOAD*HMW1_LOAD_PERCENT/mab_molecular_weight
c_hmw2 = 0.01*C_LOAD*HMW2_LOAD_PERCENT/mab_molecular_weight
c_load_array = np.array([C_SALT, c_mab, c_hmw1, c_hmw2])
""" CADET PROCESS MODEL ___________________________________________________ """
# Component System
component_system = ComponentSystem()
component_system.add_component('Salt')
component_system.add_component('mAb')
component_system.add_component('HMW 1')
component_system.add_component('HMW 2')
# Binding Model
binding_model = MultiComponentColloidal(component_system, name='Colloidal')
binding_model.is_kinetic = False
binding_model.bound_states = [0, 1, 1, 1]
binding_model.kappa_factor = 0 #k_f
binding_model.kappa_exponential = 0 #k_ef
binding_model.kappa_constant = 4.0E-9 # k_c, m
# ln Ke_i = ka_i * c0^(-kb_i) + kc_i * exp(kd_i * c0)
binding_model.logkeq_power_factor = [ 0, 3.5, 5.75, 5.5] # k_a,i
binding_model.logkeq_power_exponent = [ 0, 0, 0, 0] # k_b,i
binding_model.logkeq_exponent_factor = [ 0, 0, 0, 0] # k_c,i
binding_model.logkeq_exponent_multiplier=[ 0, 0, 0, 0] # k_d,i
binding_model.logkeq_ph_exponent = [ 0, 0, 0, 0] # set nonzero if using pH pseudo-comp
# b_pp,i = ba_i * c0^(bb_i) + bc_i * exp(bd_i * c0)
binding_model.bpp_power_factor = [ 1, 0.5, 0.5, 0.5] # b_a,i
binding_model.bpp_power_exponent = [ 0, 0, 0, 0] # b_b,i
binding_model.bpp_exponent_factor = [ 0, 0, 0, 0] # b_c,i
binding_model.bpp_exponent_multiplier = [ 0, 0, 0, 0] # b_d,i
binding_model.bpp_ph_exponent = [ 0, 0, 0, 0] # set if modeling pH dependence
binding_model.use_ph = False # True if you add a pH pseudo-component (component 1)
binding_model.phase_ratio = 4.5e7 # example; tune to your resin
binding_model.coordination_number = 6
binding_model.protein_radius = [1.59e-9, 4.5e-9, 4.5e-9, 4.5e-9]
binding_model.kinetic_rate_constant = [1e10, 1e10, 1e10, 1e10] # 1/s
binding_model.linear_threshold = 1e-7 # mol/m^3
# Unit Operations
inlet = Inlet(component_system, name='inlet')
inlet.flow_rate = FLOWRATE/(1000000*60) # m^3/s
column = GeneralRateModel(component_system, name='column')
column.binding_model = binding_model
column.length = column_length
column.diameter = column_dia
column.bed_porosity = column_porosity
column.particle_radius = particle_diameter
column.particle_porosity = particle_porosity
column.axial_dispersion = 1e-6
column.film_diffusion = column.n_comp * [1e-4]
Dp_mAb = 6.0E-12
column.pore_diffusion = [7e-10, Dp_mAb, Dp_mAb/6, Dp_mAb/12]
column.surface_diffusion = [4*50E-14, 1*7E-14, 1*7E-14]
column.c = [C_SALT, 0, 0, 0]
column.cp = [C_SALT, 0, 0, 0]
column.q = [0, 0, 0]
grm_discretization = GRMDiscretizationFV()
grm_discretization.npar = 30 # This sets an appropriate number of particle shells
grm_discretization.ncol = 50 # This sets an appropriate number of column slices
column.discretization = grm_discretization # This attaches the above discretization scheme
column.solution_recorder.write_solution_bulk = True
column.solution_recorder.write_solution_particle = True
column.solution_recorder.write_solution_solid = True
outlet = Outlet(component_system, name='outlet')
# --- New: PFR for tubing ---
pfr = LumpedRateModelWithoutPores(component_system, name='pfr_tubing')
pfr.length = 1.1 # [m] example tubing length
pfr.diameter = 0.0005 # [m] example ID
pfr.axial_dispersion = 1e-6 # small dispersion; adjust as needed
pfr.total_porosity = 0.99
""" CADET PROCESS FLOWSHEET AND RUN _______________________________________ """
# Flow Sheet
flow_sheet = FlowSheet(component_system)
flow_sheet.add_unit(inlet)
#flow_sheet.add_unit(pfr)
flow_sheet.add_unit(column)
flow_sheet.add_unit(outlet, product_outlet=True)
flow_sheet.add_connection(inlet, column)
#flow_sheet.add_connection(inlet, pfr)
#flow_sheet.add_connection(pfr, column)
flow_sheet.add_connection(column, outlet)
# Process
process = Process(flow_sheet, 'lwe')
load_duration = 60*VOL_LOAD/FLOWRATE
wash_duration = 60*COL_VOL*COLVOL_WASH/FLOWRATE
t_gradient_start = load_duration + wash_duration
gradient_duration = 60*COL_VOL*COLVOL_GRAD/FLOWRATE
process.cycle_time = load_duration + wash_duration + gradient_duration
c_load = c_load_array
c_wash = np.array([C_SALT, 0.0, 0.0, 0.0])
c_elute = np.array([500.0, 0.0, 0.0, 0.0])
gradient_slope = (c_elute - c_wash) / gradient_duration
c_gradient_poly = np.array(list(zip(c_wash, gradient_slope)))
process.add_event('load', 'flow_sheet.inlet.c', c_load)
process.add_event('wash', 'flow_sheet.inlet.c', c_wash, load_duration)
process.add_event('grad_start', 'flow_sheet.inlet.c', c_gradient_poly, t_gradient_start)
start_time = time.perf_counter()
if MODEL_CADET == True:
print("Running the chromatography model...")
if __name__ == '__main__':
from CADETProcess.simulator import Cadet
process_simulator = Cadet()
simulation_results = process_simulator.simulate(process)
end_time = time.perf_counter()
# Extract solution data in dedicated arrays
solution_time = simulation_results.solution.column.outlet.coordinates['time']
solution_mL = solution_time*inlet.flow_rate[0]*1000*1000 + DEAD_VOL #mL
solution_salt = simulation_results.solution.column.outlet.total_concentration_components[:,0]
solution_cond = solution_salt*0.103 # mS/cm
solution_mab = simulation_results.solution.column.outlet.total_concentration_components[:,1]
solution_hmw1 = simulation_results.solution.column.outlet.total_concentration_components[:,2]
solution_hmw2 = simulation_results.solution.column.outlet.total_concentration_components[:,3]
solution_total = solution_mab + solution_hmw1+solution_hmw2
ax1.plot(solution_mL, mab_molecular_weight*solution_mab, 'b--', linewidth=2)
ax1.plot(solution_mL, HMW_SCALE_FACTOR*mab_molecular_weight*solution_hmw1, 'y--', linewidth=2)
ax1.plot(solution_mL, HMW_SCALE_FACTOR*mab_molecular_weight*solution_hmw2, 'r--', linewidth=2)
ax1.plot(solution_mL, mab_molecular_weight*solution_total, 'k--', linewidth=4)
ax2.plot(solution_mL, solution_cond, linewidth=2, color = 'grey')
ax1.set_xlim(0, 25)
ax1.set_ylim(0, 15)
else:
print("Skipping to run the chromatography model. Only plots.")
# Finalize plot
plt.title("")
fig.tight_layout()
plt.show()
elapsed_time = end_time - start_time
print(f"Solver time: {elapsed_time:.4f} seconds")