Hi Everyone,
I have tried using cadet-process (being a new user to cadet hence more user friendly), but i am unable to import source and sink post installing cadet-process and am facing import error.
Below is the code used:
from CADETProcess.processModel import ComponentSystem
from CADETProcess.processModel import StericMassAction
from CADETProcess.processModel import Source, GeneralRateModel, Sink
from CADETProcess.processModel import FlowSheet
from CADETProcess.processModel import Process
Below is the Error reverted with:
ImportError: cannot import name 'Source' from 'CADETProcess.processModel' (/usr/local/lib/python3.10/dist-packages/CADETProcess/processModel/__init__.py)
PS: I am using google Colab as interface for code and have pip installed latest version of CADET-process
I am not facing error in importing any other function but only for source and sink, so if we can bypass these functions in below code, things should be fine
import numpy as np
from CADETProcess.processModel import ComponentSystem
from CADETProcess.processModel import StericMassAction
from CADETProcess.processModel import Source, GeneralRateModel, Sink
from CADETProcess.processModel import FlowSheet
from CADETProcess.processModel import Process
# Input Model
c_salt_low = 84.105
c_salt_high = 288.58
c_protein = 0.029
# Component System
component_system = ComponentSystem()
component_system.add_component('Salt')
component_system.add_component('Protein')
# Binding Model
binding_model = StericMassAction(component_system, name='SMA')
binding_model.is_kinetic = True
binding_model.adsorption_rate = [0.0, 4.35E-31]
binding_model.desorption_rate = [0.0, 1.00E-22]
binding_model.characteristic_charge = [0.0, 14.000]
binding_model.steric_factor = [0.0, 261.000]
binding_model.capacity = 710
# Unit Operations
inlet = Source(component_system, name='inlet')
inlet.flow_rate = 2.83e-9
column = GeneralRateModel(component_system, name='column')
column.binding_model = binding_model
column.length = 0.05
column.diameter = 0.15e-2*2
column.bed_porosity = 0.47
column.particle_radius = 3.25E-05
column.particle_porosity = 0.23
column.axial_dispersion = 3.9e-7
column.film_diffusion = [2.0e-5, 1.0e-5]
column.pore_diffusion = [7e-5, 5.5e-12]
column.surface_diffusion = [0.0, 1.38E-15]
column.c = [c_salt_low, 0]
column.q = [binding_model.capacity, 0]
outlet = Sink(component_system, name='outlet')
# flow sheet
flow_sheet = FlowSheet(component_system)
flow_sheet.add_unit(inlet)
flow_sheet.add_unit(column)
flow_sheet.add_unit(outlet, chromatogram_sink=True)
flow_sheet.add_connection(inlet, column)
flow_sheet.add_connection(column, outlet)
# Process
process = Process(flow_sheet, 'lwe')
process.cycle_time = 9898.902
## Create Events and Durations
wash_start = 4232.598
gradient_start = 7407.137
concentration_difference = np.array([c_salt_high, 0.0]) - np.array([c_salt_low, 0.0])
gradient_duration = process.cycle_time - gradient_start
gradient_slope = concentration_difference/gradient_duration
process.add_event('load', 'flow_sheet.inlet.c', [c_salt_low, c_protein])
process.add_event('wash', 'flow_sheet.inlet.c', [c_salt_low, 0.0], wash_start)
process.add_event(
'grad_start',
'flow_sheet.inlet.c',
[[c_salt_low, gradient_slope[0]], [0, gradient_slope[1]]],
gradient_start
)
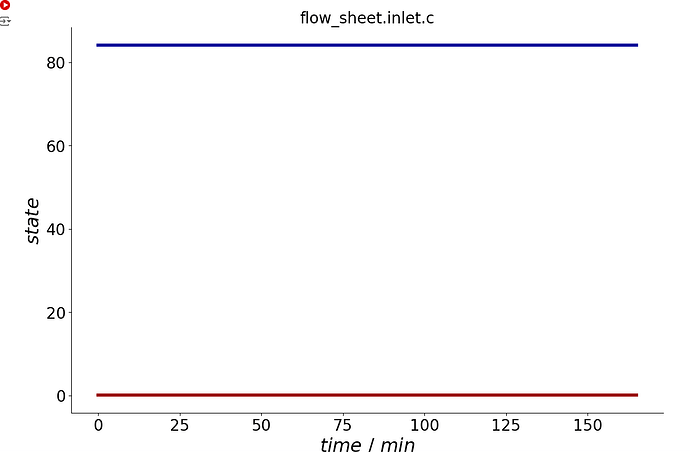
process.plot_events()
from CADETProcess.simulator import Cadet
process_simulator = Cadet()
simulation_results = process_simulator.simulate(process)
from CADETProcess.plotting import SecondaryAxis
sec = SecondaryAxis()
sec.component_indices = [0]
sec.y_label = '$c_{salt}$'
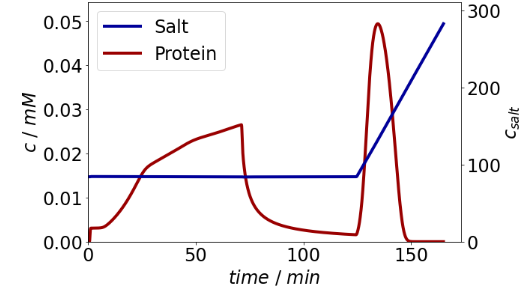
simulation_results.solution.column.outlet.plot(secondary_axis=sec)