During the workshop, I realized something odd about using the rapid equilibrium in adsorption models.
If I set the following initial conditions in a CSTR (without any in- or outgoing flow)
linear_model.root.input.model.unit_000.init_c = [1]
linear_model.root.input.model.unit_000.init_q = [0]
and set is_kinetic to True
## Adsorption
linear_model.root.input.model.unit_000.nbound = [1]
linear_model.root.input.model.unit_000.adsorption_model = 'LINEAR'
linear_model.root.input.model.unit_000.adsorption.is_kinetic = True
linear_model.root.input.model.unit_000.adsorption.lin_ka = [2]
linear_model.root.input.model.unit_000.adsorption.lin_kd = [1]
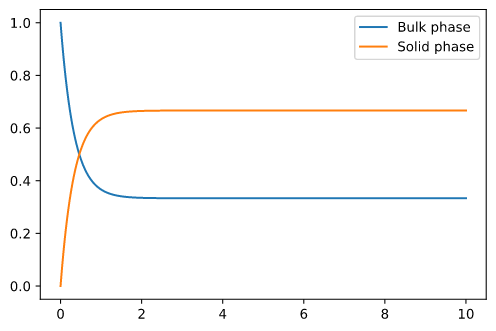
I get a graph that looks as expected:
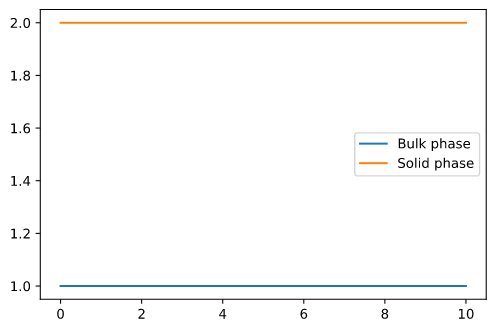
However, if i set is_kinetic = False, it seems to ignore the initial conditions. Mass is created from no where (expected would be 0.66 in solid phase and 0.33 in bulk phase).
lieres
November 10, 2020, 8:59am
2
I assume this might be caused by the consistent initialization.
This sounds like a consistent initialization problem, indeed.@j.schmoelder can you tell me the porosity to reproduce this problem?
For this example, I set the porosity to 0.5
Issue identified; Discussion continues on Github:
opened 08:11AM - 21 Oct 21 UTC
bug
BindingModel
During the workshop, we realized something odd about using the rapid equilibrium… in adsorption models.
When setting the following initial conditions in a CSTR (without any in- or outgoing flow)
```
linear_model.root.input.model.unit_000.init_c = [1]
linear_model.root.input.model.unit_000.init_q = [0]
```
and setting `is_kinetic` to `True`
```
## Adsorption
linear_model.root.input.model.unit_000.nbound = [1]
linear_model.root.input.model.unit_000.adsorption_model = 'LINEAR'
linear_model.root.input.model.unit_000.adsorption.is_kinetic = True
linear_model.root.input.model.unit_000.adsorption.lin_ka = [2]
linear_model.root.input.model.unit_000.adsorption.lin_kd = [1]
```
We get a graph that looks as expected when plotting c and q over time:

However, when setting `is_kinetic = False`, it seems to ignore the initial conditions. Mass is created from nowhere (expected would be 0.66 in solid phase and 0.33 in bulk phase):