Hello all,
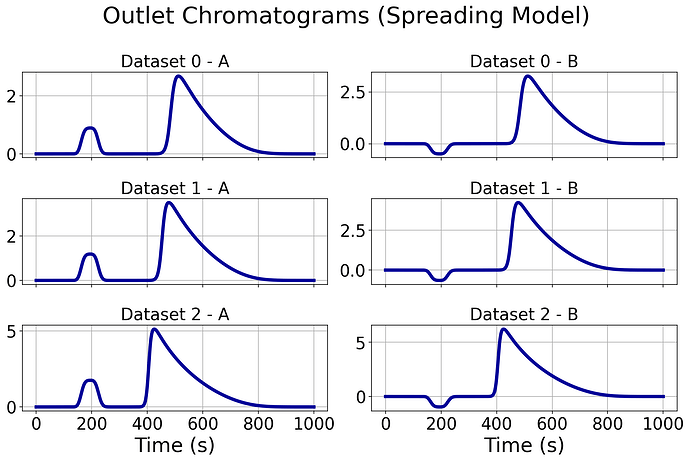
I am trying to work with the Multi-Component Spreading Binding model. My end goal is to simulate a process with 3 components (A, B, and C) however I am having issues generating results. In order to trouble-fix, I am first looking at a process with just 2 components (A and B). My main issue is I keep getting negative concentrations at the outlet, so I decided to set site exchange rate constants to 0 to allow no exchange in the hope I could replicate the results from CODE 2 which is a working Multicomponent Bi-Langmuir model, however I still see this same negative concentration issue. This attempt is shown in CODE 1.
CODE 1 - attempt to replicate CODE 2 with spreading model:
# ============================================================
# Imports
# ============================================================
import numpy as np
import matplotlib.pyplot as plt
from CADETProcess.processModel import ComponentSystem
from CADETProcess.processModel import Spreading
from CADETProcess.processModel import (
Inlet, LumpedRateModelWithoutPores, Outlet
)
from CADETProcess.processModel import FlowSheet, Process
# ============================================================
# Feed concentrations (NOW 2 COMPONENTS)
# ============================================================
C_in = [
[7.5, 7.5],
[10.0, 10.0],
[15.0, 15.0]
]
c_store = []
q_store = []
# ============================================================
# Loop over datasets
# ============================================================
for i in range(len(C_in)):
# --------------------------------------------------------
# Component system
# --------------------------------------------------------
component_system = ComponentSystem()
component_system.add_component('A')
component_system.add_component('B')
#component_system.add_component('C')
# --------------------------------------------------------
# Spreading binding model (STATE SWITCHING)
# --------------------------------------------------------
binding_model = Spreading(component_system, n_binding_sites=2, name='spreading')
binding_model.is_kinetic = True
# IMPORTANT: multiple binding states required
#binding_model.n_binding_sites = 2
# adsorption/desorption (per component)
binding_model.adsorption_rate = [
0.015, 0.03, # state 1 (A, B, C)
0.02, 0.035 # state 2 (A, B, C)
]
binding_model.desorption_rate = [
1.0, 1.0,
1.0, 1.0
]
# capacity per state
binding_model.capacity = [
100.0, 100.0, # state 1 (A, B, C)
100.0, 100.0 # state 2 (A, B, C)
]
# --------------------------------------------------------
# STATE SWITCHING (KEY FEATURE)
# --------------------------------------------------------
# switching: state 1 -> state 2
binding_model.exchange_from_1_2 = [0.0, 0.0]
# switching: state 2 -> state 1
binding_model.exchange_from_2_1 = [0.0, 0.0]
print("Spreading model configured")
# --------------------------------------------------------
# Column system
# --------------------------------------------------------
feed = Inlet(component_system, name='feed')
feed.c = C_in[i]
eluent = Inlet(component_system, name='eluent')
eluent.c = [0, 0]
column = LumpedRateModelWithoutPores(component_system, name='column')
column.binding_model = binding_model
column.length = 0.6
column.diameter = 0.024
column.axial_dispersion = 4.7e-6
column.total_porosity = 0.65
column.solution_recorder.write_solution_bulk = True
column.solution_recorder.write_solution_solid = False
column.solution_recorder.split_components = True
column.solution_recorder.write_solution_bound = True
outlet = Outlet(component_system, name='outlet')
# --------------------------------------------------------
# Flow sheet
# --------------------------------------------------------
flow_sheet = FlowSheet(component_system)
flow_sheet.add_unit(feed, feed_inlet=True)
flow_sheet.add_unit(eluent, eluent_inlet=True)
flow_sheet.add_unit(column)
flow_sheet.add_unit(outlet, product_outlet=True)
flow_sheet.add_connection(feed, column)
flow_sheet.add_connection(eluent, column)
flow_sheet.add_connection(column, outlet)
process = Process(flow_sheet, 'batch elution (Spreading)')
# --------------------------------------------------------
# Events
# --------------------------------------------------------
Q = 60 / (60 * 1e6)
process.add_event('feed_on', 'flow_sheet.feed.flow_rate', Q)
process.add_event('feed_off', 'flow_sheet.feed.flow_rate', 0.0)
process.add_event('eluent_on', 'flow_sheet.eluent.flow_rate', Q)
process.add_event('eluent_off', 'flow_sheet.eluent.flow_rate', 0.0)
process.add_duration('feed_duration')
process.add_event_dependency('eluent_on', ['feed_off'])
process.add_event_dependency('eluent_off', ['feed_on'])
process.add_event_dependency('feed_off', ['feed_on','feed_duration'], [1,1])
process.cycle_time = 1000
process.feed_duration.time = 60
# --------------------------------------------------------
# Run CADET
# --------------------------------------------------------
print('here 1')
from CADETProcess.simulator import Cadet
print('here 2')
simulator = Cadet()
print('here 3')
simulation_results = simulator.simulate(process)
print('here 4')
# --------------------------------------------------------
# Extract results
# --------------------------------------------------------
print('here 5')
outlet_solution = simulation_results.solution.column.outlet
bound_solution = simulation_results.solution.column.bound
print('here 6')
t_full = outlet_solution.time
c = outlet_solution.solution
print('here 7')
c_store.append(c)
q_store.append(bound_solution.solution)
c_store = np.array(c_store)
print("DATA GENERATED (SPREADING MODEL)\n")
# ============================================================
# Plot outlet concentrations
# ============================================================
plt.figure(figsize=(8,6))
for i in range(len(C_in)):
c = c_store[i]
plt.plot(t_full, c[:,0], label=f"A dataset {i}")
plt.plot(t_full, c[:,1], '--', label=f"B dataset {i}")
#plt.plot(t_full, c[:,2], ':', label=f"C dataset {i}")
plt.xlabel("Time (s)")
plt.ylabel("Outlet concentration")
plt.title("Outlet Chromatograms (Spreading Model)")
plt.legend()
plt.grid()
plt.show()
# ============================================================
# Subplots per component
# ============================================================
fig, ax = plt.subplots(3, 2, figsize=(12,8), sharex=True)
labels = ['A', 'B']
for i in range(len(C_in)):
c = c_store[i]
for j in range(2):
ax[i,j].plot(t_full, c[:,j])
ax[i,j].set_title(f"Dataset {i} - {labels[j]}")
ax[i,j].grid()
for j in range(2):
ax[2,j].set_xlabel("Time (s)")
plt.suptitle("Outlet Chromatograms (Spreading Model)")
plt.tight_layout()
plt.show()
CODE 1 generates:
CODE 2:
import numpy as np
import matplotlib.pyplot as plt
# ============================================================
# CADET Simulation (Generate Synthetic Truth)
# ============================================================
from CADETProcess.processModel import ComponentSystem
from CADETProcess.processModel import BiLangmuir
from CADETProcess.processModel import (
Inlet, LumpedRateModelWithoutPores, Outlet
)
from CADETProcess.processModel import FlowSheet, Process
C_in = [[7.5, 7.5], [10, 10], [15, 15]]
c_store = []
q_store = []
for i in range(len(C_in)):
component_system = ComponentSystem()
component_system.add_component('A')
component_system.add_component('B')
# --------------------------------------------------------
# Bi-Langmuir binding model
# --------------------------------------------------------
binding_model = BiLangmuir(component_system, n_binding_sites=2, name='langmuir')
binding_model.is_kinetic = True
binding_model.adsorption_rate = [0.015, 0.03, 0.02, 0.035]
binding_model.desorption_rate = [1, 1, 1, 1]
binding_model.capacity = [100, 100, 100, 100]
#binding_model.bound_states = [2, 2]##
print('hi 1')
# --------------------------------------------------------
# Column system
# --------------------------------------------------------
feed = Inlet(component_system, name='feed')
feed.c = C_in[i]
eluent = Inlet(component_system, name='eluent')
eluent.c = [0, 0]
column = LumpedRateModelWithoutPores(component_system, name='column')
column.binding_model = binding_model
column.length = 0.6
column.diameter = 0.024
column.axial_dispersion = 4.7e-6
column.total_porosity = 0.65
column.solution_recorder.write_solution_bulk = True
column.solution_recorder.write_solution_solid = False
column.solution_recorder.split_components = True
column.solution_recorder.write_solution_bound = True # records bound states per site
#column.solution_recorder.split_components = False
#column.solution_recorder.write_solution_bound = True
outlet = Outlet(component_system, name='outlet')
flow_sheet = FlowSheet(component_system)
flow_sheet.add_unit(feed, feed_inlet=True)
flow_sheet.add_unit(eluent, eluent_inlet=True)
flow_sheet.add_unit(column)
flow_sheet.add_unit(outlet, product_outlet=True)
flow_sheet.add_connection(feed, column)
flow_sheet.add_connection(eluent, column)
flow_sheet.add_connection(column, outlet)
process = Process(flow_sheet, 'batch elution')
print('hi 2')
# --------------------------------------------------------
# Events
# --------------------------------------------------------
Q = 60 / (60 * 1e6)
process.add_event('feed_on', 'flow_sheet.feed.flow_rate', Q)
process.add_event('feed_off', 'flow_sheet.feed.flow_rate', 0.0)
process.add_event('eluent_on', 'flow_sheet.eluent.flow_rate', Q)
process.add_event('eluent_off', 'flow_sheet.eluent.flow_rate', 0.0)
process.add_duration('feed_duration')
process.add_event_dependency('eluent_on', ['feed_off'])
process.add_event_dependency('eluent_off', ['feed_on'])
process.add_event_dependency('feed_off', ['feed_on','feed_duration'], [1,1])
process.cycle_time = 1000
process.feed_duration.time = 60
print('hi 3')
# --------------------------------------------------------
# Run CADET
# --------------------------------------------------------
print(binding_model.n_binding_sites)
print(binding_model.n_bound_states)
if __name__ == "__main__":
from CADETProcess.simulator import Cadet
print('hi 4')
process_simulator = Cadet()
print('hi 5')
simulation_results = process_simulator.simulate(process)
print('hi 6')
# --------------------------------------------------------
# Extract outlet solution
# --------------------------------------------------------
outlet_solution = simulation_results.solution.column.outlet
solid_solution = simulation_results.solution.column.solid
bound_solution = simulation_results.solution.column.bound
q = bound_solution.solution
#q_store.append(q)
t_full = outlet_solution.time
c = outlet_solution.solution
c_store.append(c)
bound_solution = simulation_results.solution.column.bound
q_store.append(bound_solution.solution)
c_store = np.array(c_store)
#q_store = np.array(q_store)
print("DATA GENERATED\n")
# ============================================================
# Plot outlet concentration
# ============================================================
plt.figure(figsize=(8,6))
for i in range(len(C_in)):
c = c_store[i]
cA = c[:,0]
cB = c[:,1]
plt.plot(t_full, cA, label=f"A dataset {i}")
plt.plot(t_full, cB, '--', label=f"B dataset {i}")
plt.xlabel("Time (s)")
plt.ylabel("Outlet concentration")
plt.title("Outlet Chromatograms (Bi-Langmuir)")
plt.legend()
plt.grid()
plt.show()
# ============================================================
# Plot outlet concentration
# ============================================================
fig, ax = plt.subplots(3, 2, figsize=(10,8), sharex=True)
for i in range(len(C_in)):
c = c_store[i]
cA = c[:,0]
cB = c[:,1]
# A plots (left column)
ax[i,0].plot(t_full, cA)
ax[i,0].set_title(f"Dataset {i} - A")
ax[i,0].set_ylabel("Concentration")
ax[i,0].grid()
# B plots (right column)
ax[i,1].plot(t_full, cB)
ax[i,1].set_title(f"Dataset {i} - B")
ax[i,1].grid()
# x labels only on bottom row
ax[2,0].set_xlabel("Time (s)")
ax[2,1].set_xlabel("Time (s)")
plt.suptitle("Outlet Chromatograms (Bi-Langmuir)")
plt.tight_layout()
plt.show()
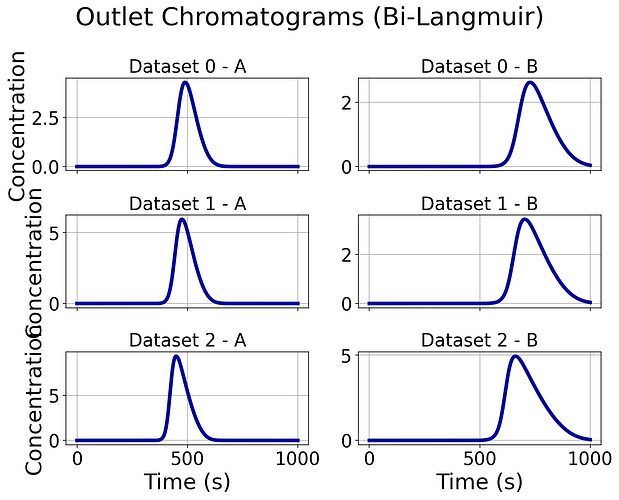
CODE 2 generates:
Am I setting something up wrong that may cause these negative concentrations?
Many thanks!